Awards and recognition
Seed funding yields high rewards for 2019 Kickstart Catalyst Award winners
Written by MSL Postdoc Communications Team including Andreas Geissner and Zack Gerbec.
With research ranging from characterizing tree defense mechanisms to identifying calcium channel isoforms, the Michael Smith Laboratories is truly a multidisciplinary unit. Established in 2017 as the ‘Director’s Kickstart Award’, the competition which was recently renamed to “Kickstart Catalyst Award”, aims to fund and develop unique and different collaborations, giving researchers a new perspective to their research questions. This award empowers graduate students, postdoctoral fellows, and faculty members to attempt high-risk and high-reward projects. The 2019 awardees shared their preliminary findings on October 23rd 2019, in the “Kickstart Project Presentations”. These awards support collaborative initiatives aimed at the generation of preliminary data for future grant applications to external funding agencies.
Below are graphical abstracts and project summaries of the 2019 award recipients:
Conifer secretory trichomes – characterizing the transcriptome, proteome, and metabolome of a new type of specialized cell factories for conifer defense
Awardees: Jose Celedon, Research Associate in the Bohlmann Lab, and Dr. Leonard Foster, Principal Investigator
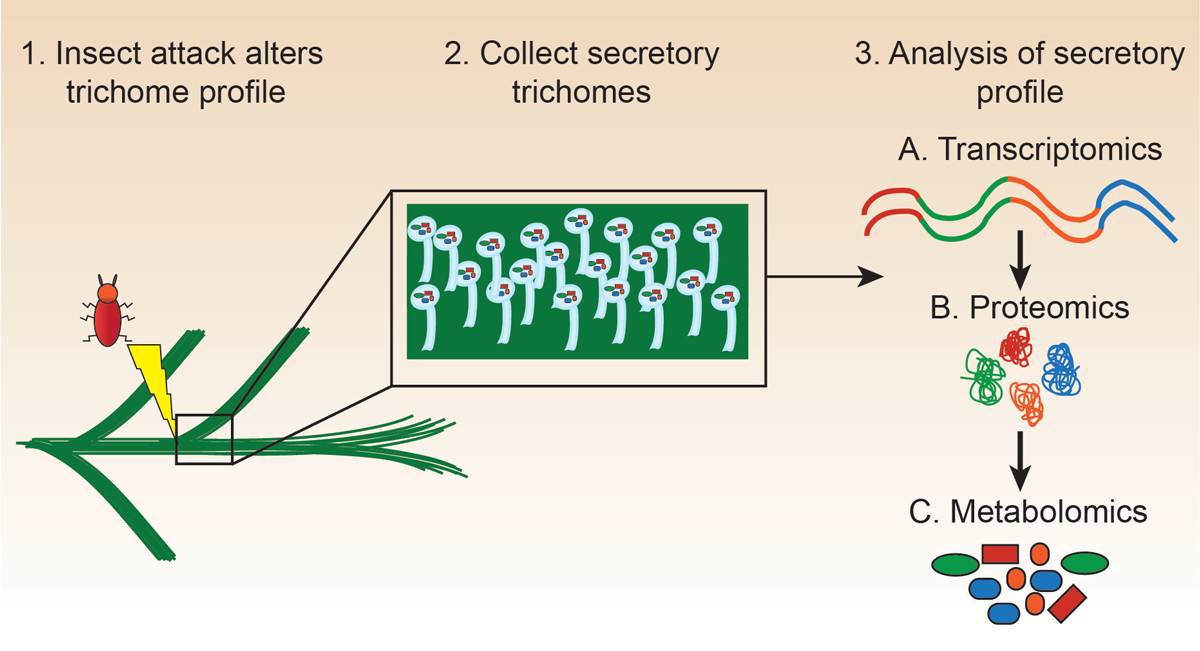
Coniferous forests represent a critical natural resource that provides valuable material goods such as chemicals and paper products and contributes substantially to ecosystem homeostasis through carbon sequestration. As part of the Kickstart Award, the Foster and Bohlmann labs sought to understand the defense mechanisms present in coniferous plants that enable them to adapt and survive in changing environments.
The labs utilized the Kickstart funding to characterize the molecular contents of trichomes collected from conifers grown in the lab. These trichomes are molecular nano-factories that produce a wide array of compounds that facilitate interactions with the plant’s environment. These interactions enable plants to defend themselves against insects and to provide protection from other pathogens. Using the combined expertise of both labs, the researchers were able to define the transcriptome, metabolome, and proteome present in coniferous trichomes. While the Bohlmann lab is well-versed in transcriptomic and metabolomics analyses, collecting the proteomic data would not have been possible without contributions from the Foster lab.
“Together, all three approaches provided a complete picture of trichomes biosynthetic machinery to explain the chemical diversity of their metabolomes,” says Jose describing the project in an interview. Through this collaborative effort, researchers were able to identify several novel metabolites and the potential proteins responsible for producing them. Further analyses of the results produced from this Kickstart collaboration have the potential to improve conifer breeding and sustainability by optimizing plant defense against insects and pathogens.
Manipulating primary Type 2 innate lymphocytes (ILC2) in vitro
Awardees: Pablo De Lucia Finkel, a Graduate Student in the Jefferies Lab, and Dr. Wilfred Jefferies, Principal Investigator, and Dr. James Piret, Principal Investigator
While chemotherapy continues to be a critical part of cancer treatment, the significant side effects associated with chemotherapeutic agents have lead researchers to explore other options. One such alternative termed ‘immunotherapy,’ where the immune system is utilized to directly target cancerous cells, has demonstrated clinical efficacy and has the potential to provide better specificity than many forms of chemotherapy. A recent collaboration between the Jefferies and Piret labs is attempting to expand upon current immunotherapy methods by bringing a new tool to the immunotherapy toolbox.
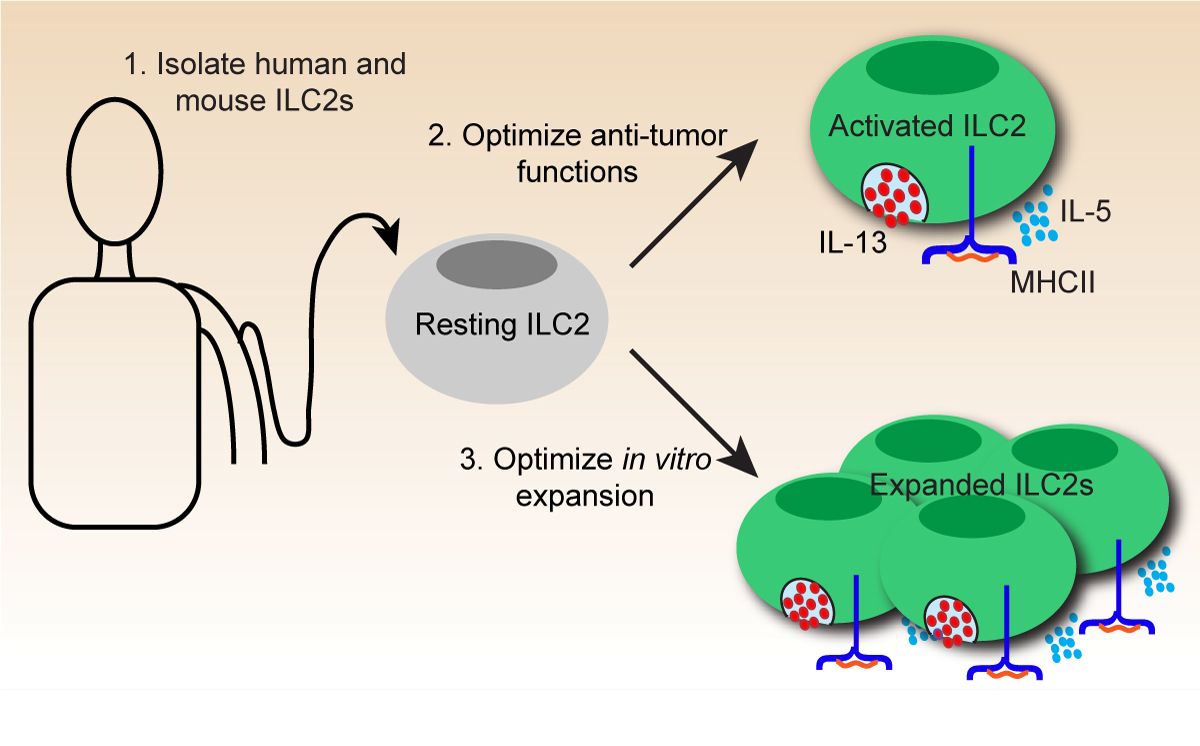
In cell-based immunotherapy, immune cells are functionally optimized to target and eliminate cancer cells prior to being transferred into patients. Using the funding provided by the Kickstart Award, the Jefferies and Piret labs explored new ways to culture and expand the largely uncharacterized immune cell Type 2 innate lymphocytes (ILC2s) in a way that would boost the anti-tumor functions of these cells. “The Piret lab is all about cell culture…and they have the expertise to do that, and we [the Jefferies lab] provide the immunology and the cell biology,” says Pablo de Lucia Finkel, a graduate student in the Jefferies Lab, when describing the collaboration. Through their group effort, the labs discovered a new method for expanding ILC2s in culture; a critical step in optimizing immunotherapy.
While the hope is that these cells can be used for cancer treatment one day, there is also an immediate, basic science benefit from this work as an improved ability to expand these cells in culture will make them easier to study. These labs hope to follow up on the preliminary findings to not only improve cancer treatment, but to contribute to basic science knowledge that forms the foundation of biomedical research.
Selective disruption of bacterial communication using Quorum Quenching enzymes to modulate microbiota composition
Awardees: Jorge Peña Díaz, Graduate Student in the Finlay Lab, and Rachel Jun, Graduate Student in the Tokuriki Lab
In the bacterial world, intra and interspecies interactions are critical for shaping the overall microbial community and determining the microbiome contribution to health and homeostasis. With the help of the Kickstart Award, Rachel and Jorge attempted to provide a better understanding of how bacteria communicate with members of their own species and how this communication affects overall group dynamics.
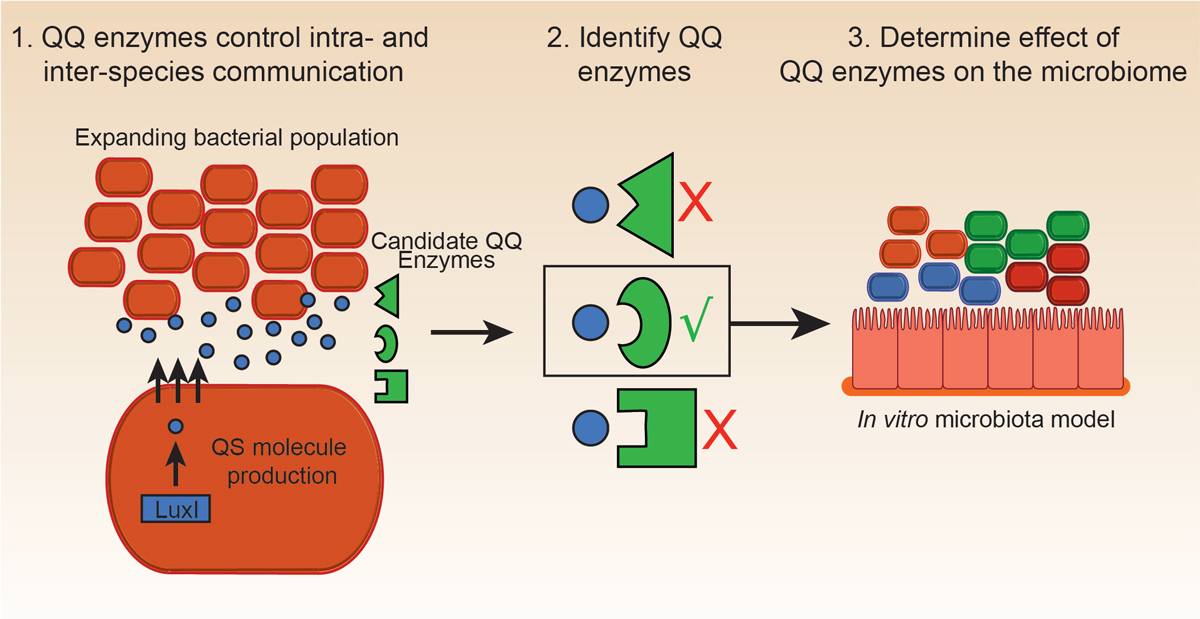
In a diverse bacterial community, members of a given species communicate through Quorum Sensing wherein specific signaling molecules are produced as bacterial populations expand. Using a laboratory model of the gut microbiome developed by Jorge, and candidate enzymes identified and purified by Rachel, these two were able to directly test the ability of specific proteins to degrade quorum signaling molecules and subsequently regulate overall microbiota composition. Jorge expressed his excitement for the collaboration saying, “There are very few opportunities in science to take a risk on a crazy project idea and follow it through.” Along with providing a unique experimental opportunity, the project also yielded promising results as the team was able to identify enzyme functions that only affected the bacterial population as a whole and not individual species in isolation experiments.
These findings demonstrate the utility of the laboratory model of the gut microbiome and the therapeutic potential of enzymes capable of regulating quorum signaling. “In the future, I would love to see these enzymes used in personalized medicine to target specific bacteria and modulate the microbiota towards a health-associated state,” says Jorge.
Identification and functional analysis of novel brain calcium channel isoforms with long-read sequencing
Awardees: Shamsuddin Bhuiyan, Graduate Student in the Pavlidis Lab, and John Tyson, Research Associate in the Snutch Lab
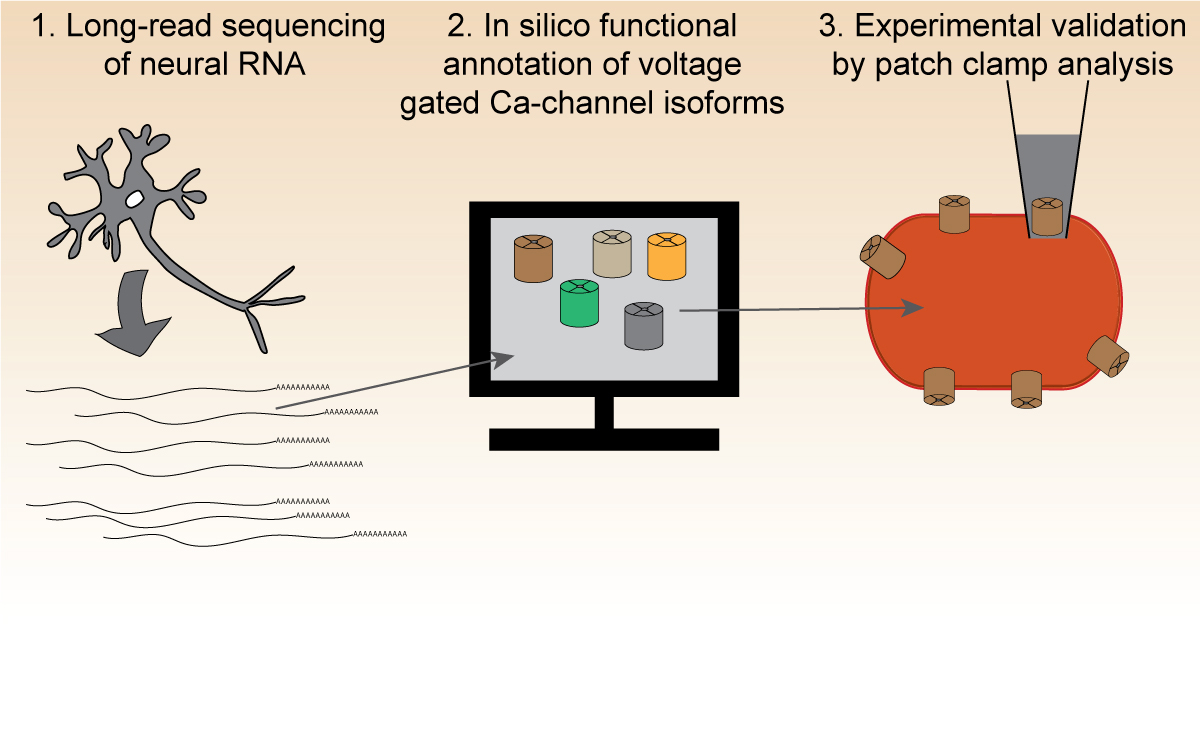
Voltage-gated calcium channels are membrane proteins that are essential for nervous system function. They are known to exist in multiple isoforms, but their variability might be more extensive than previously thought calling for an investigation into the identification and biological characterization of new isoforms.
“This was the perfect setting to use the Kickstart Award to combine the expertise of two labs that study neurodevelopmental process from a genetic perspective,” explains Shamsuddin. “The idea of the project just made sense.” The project proposed to identify and characterize new calcium channel isoforms using long-read RNA sequencing via expertise from the Snutch lab and the computational and statistical tools from the Pavlidis lab.
During the funding period, the researchers focused on one calcium channel gene termed Cav2.3, and its isoforms, and produced exciting results. “Some novel isoforms might have a different mode of regulation than previously known ones,” says Shamsuddin, for whom the study forms part of his PhD thesis. In the long term, the scientists hope to expand the research to other calcium channels and pinpoint specific splice isoforms that contribute to epilepsy.
GPCR-targeting therapeutic Abs with specific agonistic effects
Awardees: Jennifer Bui, Research Associate in the Gsponer Lab, and Roza Bidshari, Postdoctoral Fellow in the Hansen Lab/Scientist at AbCellera
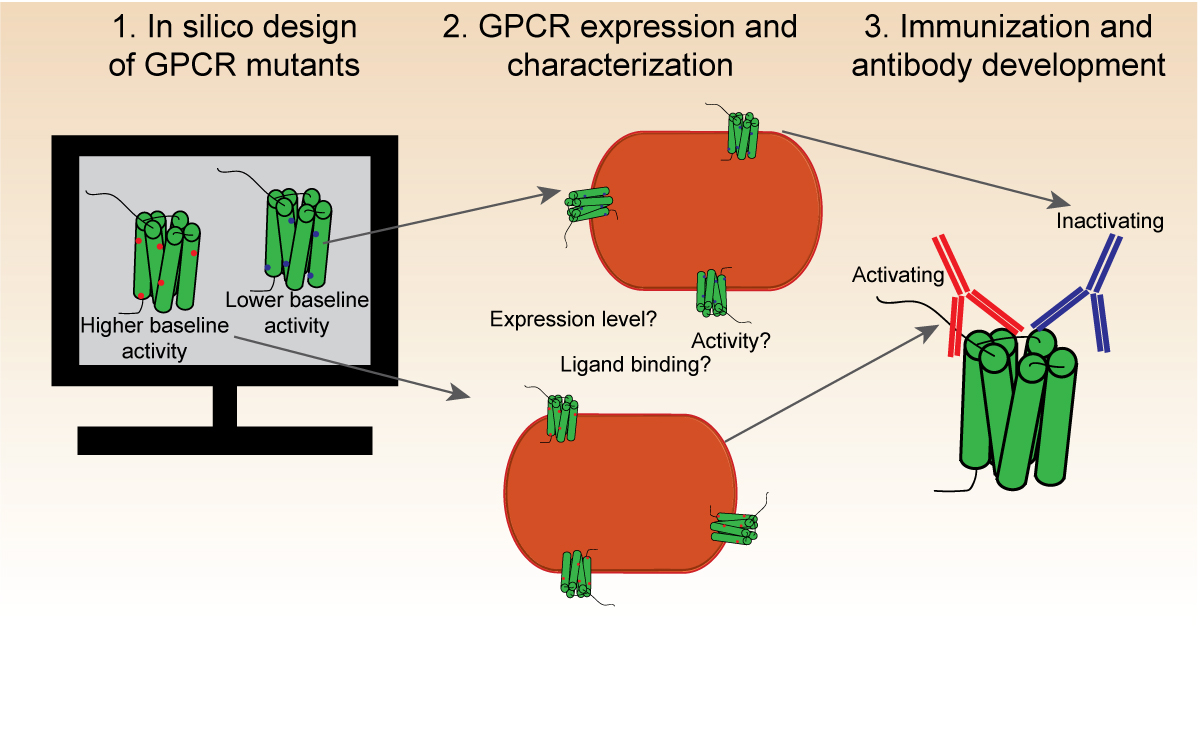
G protein-coupled receptors are proteins in the cell membranes that are switched on and off depending on the presence or absence of certain molecules nearby allowing the cell to react. Antibodies that specifically bind to the ‘on’ state could be used as an alternative way to activate G protein-coupled receptors to treat diseases
Using the Kickstart Award, the Gsponer lab pooled their expertise of computational modeling and protein design with the Hansen lab/AbCellera’s skillset of high-throughput antibody discovery to take on this challenge and isolate antibodies with this ability. The team’s approach was to immunize the G protein-coupled receptors frozen in the “on” state and then screen for antibodies from this immune response.
“It’s an ideal collaboration between local biotech and academia,” says Jennifer, who performed the computational search for new protein variants and is excited about the results they have already obtained for the first receptor they examined. “The de novo design recapitulates variants known to stabilize GPCRs and some even mimic allosteric inhibitors”.
For the future, she is thrilled about the possibilities this workflow offers once it is established, adding “we’re very excited that our established protocol could be used for therapeutic antibody discovery.”