Newsroom
The Future of Genomics in a Post Pandemic World
Over a year has passed since the pandemic hit Canada and the lockdowns began. While businesses, schools, institutions, public and health services were forced to shut down or reinvent themselves, so too were scientific research laboratories. Live experiments, grant funded research and educational activities ground to a halt as scientists around the world refocused their efforts on tackling the COVID-19 pandemic sweeping the globe. Dr. Terry Snutch of the Michael Smith Laboratories is one of many researchers at UBC that did just that. Normally working with animal models focused on epilepsy, migraine and other brain disorders, Dr. Snutch decided to repurpose the use of the Oxford Nanopore technology a portable DNA sequencer, for use decoding the COVID-19 virus. Seeing the need for a coordinated approach to tackling the emerging public health crisis in Canada, Dr. Snutch rapidly reached out to public health labs and sequencing centres across the country to build up an organized network. With robust provincial support for the project and an injection of $40 million dollars from the federal government, the Canadian COVID Genomics Network (CanCOGeN) was up and running. Half of the network (VirusSeq) quickly got to work with the aim of sequencing up to 150,000 genomes of the SARS-CoV-2 virus while the other half (HostSeq) committed to sequencing up to 10,000 patient genomes in order to build a “biobank” of data. This crucial work has informed public health policy and decision making across the country and continues to track community spread and COVID-19 variants as they emerge, as well as inform vaccine development and breakthrough infections.
We caught up with Dr. Snutch to find out how the CanCOGeN network has grown over the last 16 months, to reflect on the notable contributions of UBC researchers to the global health crisis, to understand how the experience of the pandemic has impacted the field of genomics and to learn more about what scientists in the community are looking forward to as restrictions ease here in British Columbia.
How does it feel to be over a year into this research and the CanCOGeN network?
“Exhausting (with a laugh)! The genome variant information VirusSeq derived is shared as quickly as possible with governments and health authorities and it’s used for public policy. This timely information is why we have restrictions at airports and borders, why certain public health measure have been put in place. A lot these decisions are based on the variant data that showing there have been outbreaks, how they spreading through the community, via travel etc. At the beginning there was a considerable lag between the results of the testing and the information being shared. But now the system is working broadly across the country and the information can be shared much more rapidly.”
What was the impetus for getting this project rolling?
“We really needed to have a network across the country that could identify the variants and then use that information in meaningful ways. Without the development of VirusSeq, everyone would be doing different things, employing different strategies, and not talking to each other to best inform overall Canadian policy. It was imperative that to do this together as a network and share the information.”
“It’s one of these things where you have an idea and you wake up one day and think, you know, we should do this more broadly. Then you jump into it totally naively not having done this before. I’m not an epidemiologist, I’m not a virologist, I’m a neurobiologist. But I realized that we needed to do this nationally and I understood the technologies that would be best suited for this effort.”
How might we take the lessons we’ve learned over the past year living through the pandemic and apply them to the field of genomics, improvements for healthcare and society as a whole?
“With the SARS outbreak back in 2002 it was relative small and was contained quite quickly. After that there were a number of studies and reports generated as to how Canada should prepare for future pandemics. Unfortunately, those reports were not followed up on and there was no national plan put in place about how to react to the next pandemic.
What I hope will come out of this is that there is a permanent pandemic readiness network established. The key will be to have this network set up and funded in perpetuity. It just has be there ready to go. We can’t afford the timing, organizational and funding holdups required to start from scratch again. The network needs to be sitting there with the button ready to be pushed.”
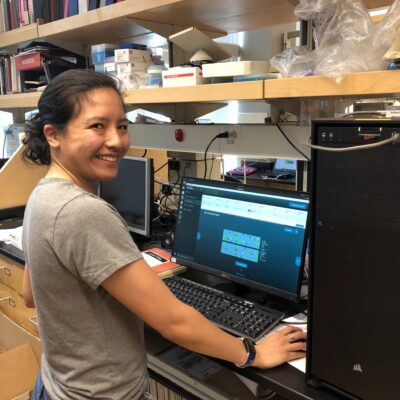

Snutch lab tech, Gabrielle Jayme, running the MinION sequencer
How has the global experience of the pandemic impacted your research area?
“It’s made the notion of the overall usefulness of genomics more widely and prominently known across the country. People now understand that genome sequencing can be used for more than just accumulating large amounts of data destined for databases, that it can be critical to the health of Canadians.
When the variants of concern started becoming headline news, many people first became aware that the virus isn’t a single virus, that there can be thousands of mutations and there was a broader recognition in the press and with politicians that we are not trying to deal with just a single virus – that it’s a moving target. We realized that unless we get to a certain level of immunity across the population nationally and globally, the virus will continue mutating, new variants will keep appearing and they will continue to be a concern if they can outmaneuver the immune response from vaccines and drug treatments. Since then there has been a real appreciation for genomics as a whole and how it can be used in both real time and towards adaptable health policy and research planning critical in regards to the pandemic.
screen shot video of a MinION run as it is sequencing in real time
Moving forward, we need a surveillance network routinely watching for these mutations and other viruses that could turn into the next pandemic, rather than wait for the next pandemic to blindside us. This should be part of a very proactive, post pandemic network across the country.”
What is the most significant finding with the genomics research?
“The biggest surprise to me was how many mutations have arisen in this virus. There are thousands and thousands already. Epidemiologists and modellers originally thought it would only mutate every 10 days to two weeks or so. But that didn’t really take into consideration that there would be 190 million people world-wide that would be infected. So there’s 190 million opportunities for those mutation cycles to happen. It is not a slow evolving virus with this level of host infections worldwide.”
How has this pandemic impacted your personal research?
“Well, our active brain research generally ground to a halt. We need staff, trainees and resources to maintain our animal colonies to keep up our experiments. All of that had to shut down and our colonies shrank dramatically. Now, we have to refocus time and money on building up those colonies and getting that research going again. We’ve lost about a year and half of progress. The whole world has been affected in this way. That’s going to affect a lot of careers as well, students graduating with their PhDs, postdocs looking for jobs. It’s very difficult to apply for jobs when labs aren’t up to speed with their own resources and research, let alone hiring new people or taking on new trainees. There is going to be gap and I worry the younger trainees will be affected the most by this.”
What is the most significant contribution from the UBC research community towards the global efforts to combat COVID-19?
“The most notable positive outcome is the many labs that were able to take advantage of the reduction in wet lab research activities and pivot to pandemic related research. To me that’s the most important thing across the entire country, that people have been able to apply their techniques and knowledge to contribute towards these pandemic efforts. It’s been a huge wake-up call that many researchers toil away in silos, rarely reaching out across disciplines. The federal government and funding agencies need to look at this and realize we couldn’t have done this without the basic research of epidemiologists, virologists, molecular biologists, modellers, bioinformaticians…everybody contributed something. And that very little of that was aimed at SARS or other types of pandemic research at the beginning. The basic research really needs to happen continually and you need to fund it well in the background. Because as we’ve all come to realize you’re going to need it at some point.”
What are you most looking forward to as we ease out of the public health restrictions?
“I look forward to taking on more trainees as soon as it is safe to do so. Working at limited capacity in the lab this past year has been challenging. I’m also looking forward to getting on airplanes going to see some of scientific colleagues in person around the world. One day I’ll count up the zoom calls I’ve done. I’ve got some great collaborations I’d like to get back to in person. It’s nice for myself and my trainees to go to a lab and actually see what they’re doing physically, talk to them one on one, go out for beers to brain storm and so on. I’m really looking forward to getting back to interacting with people in the flesh.”
What direction is your lab headed in now?
“In part, we are working on some ideas for new drugs that could be used to treat SARS-CoV-2 or similar pathogens in the future. I’m also following up on some grants for drug discovery funding. On the other hand, I’ve had a lot of ideas with regard to some new research we are doing in chronic pain and sudden unexpected death related to epilepsy. I want to apply for grants and hopefully get this research funded. Running the VirusSeq network the past 15 months has been close to seven days a week. I haven’t had a lot of time to write grants. I hope to be able to catch up.”
Thanks to the efforts of Dr. Snutch and other scientists like him, we are finally looking towards a hopeful future here in BC and Canada as whole. As infection rates and hospitalizations continue to decline across the country, the light at the end of the pandemic tunnel glows brighter. Many researchers are breathing a sigh of relief that they may begin resuming important work on the myriad of other health and environmental issues that were put aside during this critical last year. Hopefully the lessons learned from this global experience of the pandemic and the impact they have had on the field of genomics won’t be forgotten.
